Toxoplasma gondii, one of the most successful intracellular protozoan parasites, can infect the majority of vertebrate spices including humans with a worldwide distribution [
1-
3], and approximately one-third of the population has been exposed to
T. gondii. Normally, the infections are asymptomatic or subclinical. However, the
T. gondii infection can cause abortion and stillbirth in pregnant women, and encephalitis, chorioretinitis, and systemic infections in immunocompromised individuals [
2]. In animals,
T. gondii can also cause abortion in livestock, especially in sheep and goats, which can spawn a great number of economic losses in livestock [
3]. However, there was no effective vaccine and drugs that can help to control toxoplasmosis.
The strains of
T. gondii that predominate in Europe and North America, classified into types I, II, and III, differ in a wide range of phenotypes, including virulence, persistence, migratory capacity, and how they interface with the immune response [
4-
6], Thus, the information of genetic diversity of
T. gondii is useful for better understanding epidemiological patterns and pathogenicity, as well as exploring of new polymorphic virulence effectors.
GRA20, a novel dense granule protein, is secreted and targeted to parasitophorous vacuole membrane (PVM), which may participate in the manipulation of the host immunity [
7]. Previous studies have identified the existence of polymorphisms in dense granule proteins, such as GRA15, GRA5, and GRA6 [
8-
10], but the sequence variation about the
GRA20 gene among different
T. gondii isolates is still unknown. Therefore, the objective of this study was to examine sequence diversity of
GRA20 gene among
T. gondii strains from different hosts and geographical regions worldwide.
In this study, a total 16
T. gondii strains from different hosts and geographic locations were used for analysis (
Table 1). These
T. gondii isolates have been genotyped and genomic DNA (gDNA) was prepared as described previously [
11-
13].
To acquire amplicons of GRA20 genes concerning different T. gondii isolates, the primers GRA20-F (5´- ATGCATAGCCGGAACTGCGTC-3´) and GRA20-R (5´- TCACGCGGGCTTTCTACGG-3´) were designed based on T. gondii ME49 strain available in ToxoDB database (TGME49_200010). All the PCR products of GRA20 genes were purified by the DNA purification kit (GenStar, Beijing, China), ligated into pMD18-T vector (TaKaRa, Dalian, China), and then transformed into JM109 competent cells (Promega, Madison, Wisconsin, USA). Subsequently, the positive colonies were screened by PCR, and then sequenced by GenScript Co., Ltd. (Nanjing, China).
The acquired
GRA20 gene sequences were aligned by the Multiple Sequence Alignment Program, Clustal X 1.83 [
14], and sequence variation was determined among the examined
T. gondii strains. Phylogenetic reconstructions based on the complete sequences of
GRA20 gene among 13
T. gondii isolates and plus the corresponding sequences of strains TgCatBr9, VEG, and ME49 available in ToxoDB (
http://toxodb.org/toxo/) were carried out by 2 inference methods, maximum likelihood (ML) and maximum parsimony (MP) methods by Paup, with the sequence of
Hammondia hammondi (XM_008890671.1) as the out-group. Phylograms were drawn by the Tree View program version 1.65.
In the present study, the obtained entire genomic sequences of the GRA20 gene were 1,586 bp in length in all examined isolates. According to the analysis of all the 16 GRA20 complete sequences, there were 2 extrons and 1 intron in the
GRA20 gene (
Table 2). The A+T content ranged from 45.0% to 45.4% in the entire sequence. There were 124 nucleotide position variations with a distribution of 57 transitions (A↔G and C↔T), 61 transversions (C↔G, T↔G, A↔C, and A↔T) in CDS, and 2 transitions (A↔G and C↔T), and 1 transversion (C↔G, T↔G, A↔C, and A↔T) in the intron (
Table 2). However, when we analyzed the
GRA20 sequences without type III (CTG, VEG) strains, there were 12 nucleotide variations with a distribution of 10 transitions (A↔G and C↔T), 2 transversions (C↔G, T↔G, A↔C, and A↔T) in CDS, and 2 transitions (A↔G and C↔T) and 1 transversion (C↔G, T↔G, A↔C, and A↔T) in the intron. The alignment of
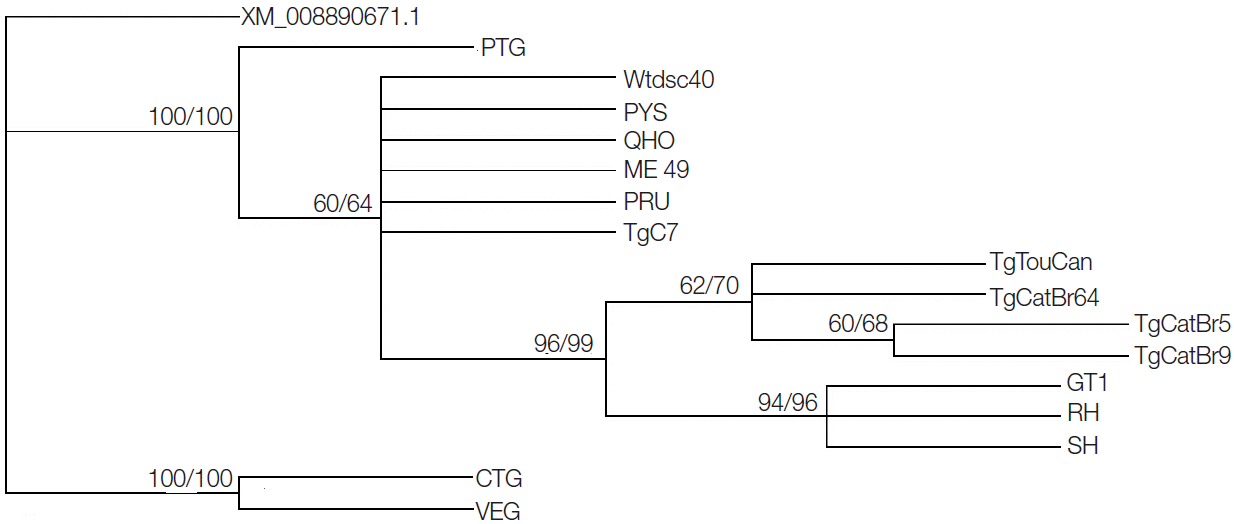
GRA20 gene sequences showed that sequence variation was 0-7.9% in all studied strains, while the sequence variation became 0-0.7% without the CTG and VEG strains. Phylogenetic reconstruction of all 16
T. gondii strains based on
GRA20 sequence data showed that the type I and type III of
T. gondii strains were clustered into respective clusters separately (
Fig. 1).
Recently, polymorphisms in the sequences of
GRA5, GRA6, GRA7, and
GRA15 genes have been reported [
8,
9,
15,
16]. Among them, polymorphic dense granule proteins were widely used in typing
T. gondii isolates, such as GRA6 [
10]. Furthermore, polymorphic dense granule protein may have different roles in regulating the inflammatory response. For example, GRA15 in type II activate more IL-12 than type I or type III strains [
8]. In this study, we found
GRA20 gene was very diverse in type III, indicating the functions may be different, too. Our results were consistent with that of some previous studies using other genetic markers, such as
GRA5, Rop17, and
HSP60 for genotyping [
9,
17,
18], but different to some previous studies, such as
Rop38 and
eIF4A [
19,
20].
In conclusion, the present study examined the sequences of the T. gondii GRA20 gene and revealed that it was more divergence in type III compared to other T. gondii strains, suggesting the functions of GRA20 in type III may be different from other strains. Phylogenetic analysis indicated that the GRA20 gene could distinguish the type I and type III strains, suggesting the GRA20 gene may be a novel genetic marker for studying genetic variation or the population genetic structures of T. gondii isolates.