Introduction
Bioindicators are organisms that indicate the quality of their environment [
1]. Thus, the environmental conditions of an area can be inferred by examining the status of these bioindicators. Inaccurate results may be obtained when performing direct measurements of the water quality due to factors such as the condition of the water and the weather on the day of measurement; the use of bioindicators can contribute to eliminating such inaccuracies [
2].
In Korea, the quality of water has been classified into 7 grades based on the Korean-Comprehensive Water Quality Index (or in terms of the biochemical oxygen demand): Grade Ia, wherein the water is considered suitable for domestic use with simple purification, and aquatic organisms such as scuds and crayfish can thrive in water of this quality; Grade Ib, wherein the water is considered suitable for domestic use following standard purification; Grade II, wherein the water can be used for swimming after standard purification (
Semisulcospira and
Plecoglossus are found in waters with these 2 grades); Grade V water is deemed normal and can be used for industrial purposes following standard purification (
Zacco platypus and specied in the genus
Pseudorasbora can be found in aquatic environments with water of this quality); Grade IV water is considered fairly poor and can be used as industrial water after having undergone advanced purification (species such as
Misgurnus spp. and
Silurus asotus are found in environments with water of this standard); Grade V is indicative of high levels of pollution, and these waters tend to be populated with more tolerant invertebrate species, such as those in the family Chironomidae and
Tubifex tubifex; and Grade VI, which has very poor quality and is unsuitable for most fish species [
3].
FLA tend to occur ubiquitously in a diverse range of terrestrial and aquatic habitats [
4], and some species, such as testate amoebae, have been used as bioindicators that are highly responsive to changes in environmental conditions. Species of testate amoebae are found in diverse environments, and their presence or absence in a particular environment can facilitate the rapid detection of environmental change. The dependence of these amoebae on specific ecosystems can indicate the health and stability of a particular ecosystem, thereby serving as a valuable diversity-based monitoring system [
5–
7].
In Korea, species of FLA in the genera
Acanthamoeba,
Vannella,
Naegleria, and
Vermamoeba are readily found in freshwater habitats [
8–
11] and are of particular medical importance. For example, the
Acanthamoeba species, which can exist as cysts or trophozoites, have been identified as causal agents of amoebic keratitis and granulomatous amoebic encephalitis [
12]. Alternatively, cysts of the
Vannella species have a fan-shaped morphology, and the trophozoites can change to a stellate form [
13,
14]. The
Naegleria species, which inhabit both soil and water, are ameboflagellate [
15] and can occur in one of 3 forms: cyst, trophozoite, and flagellate.
Naegleria fowleri has been identified as the causal organism of primary amoebic meningoencephalitis [
16].
Vermamoeba vermiformis, characterized by small round double-walled cysts and worm-like trophozoites, causes corneal inflammation and has also been reported to be associated with certain pathogenic bacteria, including
Legionella pneumophila [
17].
The aim of this study was to determine the potential use of FLA commonly found in Korean rivers as bioindicators by collecting water samples from aquatic habitats and determining the types and amounts of isolated amoebae.
Materials and Methods
Water samples
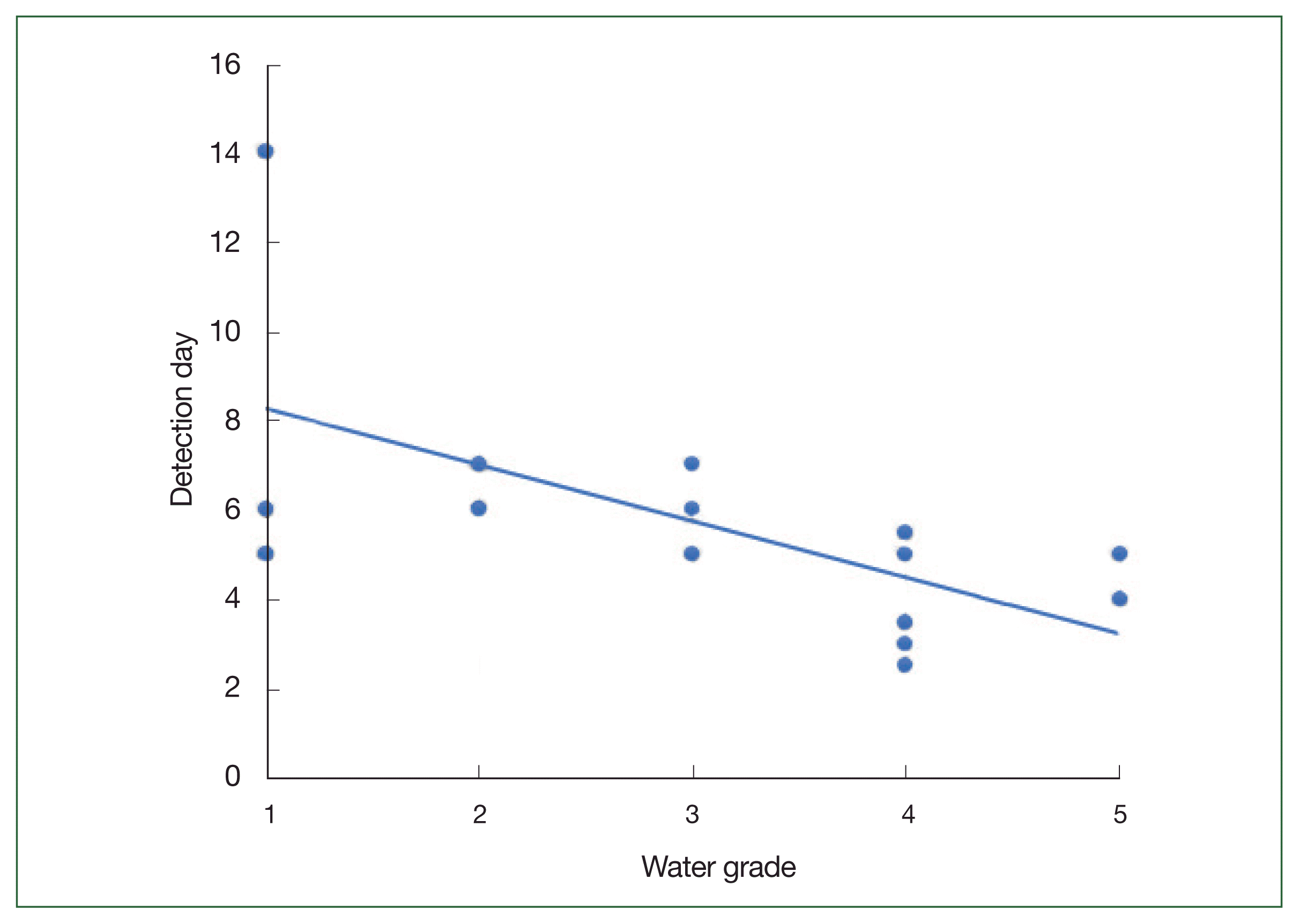
Water samples from multiple sites along the Nakdong River in Korea were collected in sterilized 1-L bottles (Winners, Busan, Korea). Each site was characterized by a specific water grade (
Fig. 1): Grade I (Ia and Ib)–Daecheon, Cheolma, Hoedong, Baenaegol, and Yusan; Grade II–Geumho, Samnak, Yangsan, and Uksu; Grade III–Mulgeum, Seonakdong, and Suyeong; Grade IV–Pyeonggang, Seokdae, Jangnim, Nakdong, and Samrak (wetland); and Grade V (V and VI)–Hwamyung and Gamjeon [
18,
19].
Preparation of agar plates
Agar medium for culturing the amoebae was prepared by initially autoclaving a 2% suspension of Bacto-agar (Biosciences, San Jose, CA, USA), which was cooled to a temperature of approximately 50°C–60°C and supplemented with amphotericin B solution (Sigma, St. Louis, MO, USA) to inhibit fungal growth. Agar plates were prepared by pouring 12 ml aliquots of the agar into Petri dishes (SPL, Pocheon, Korea); the plates were smeared with a 20% suspension of heated
Escherichia coli [
20].
Water sample filtration
Filtering apparatus (filter flask, glass base, graduated funnel, and aluminum clamp) that had initially been autoclaved and sterilized were used to filter the collected water samples. The water samples were filtered through 0.45-μm pore filters (Advantec membrane filters, Tokyo, Japan). The filters were lifted from the device using a scalpel, and forceps were inverted and placed on the surfaces of the E. coli-inoculated agar plates, which were then incubated at 25°C. The filtering apparatus was resterilized between the filtering of each sample to prevent cross-contamination.
Monoxenic and axenic cultures
Fungal growth and amoebae were microscopically checked every day after incubation. The amoebae were isolated, if detected. Regions on plates with dense amoebal growth were marked with a pen, excised, and placed on the surfaces of fresh agar plates. After obtaining the monoxenic cultures, we excised gel segments with good amoebal growth and placed them in a solution of 1% HCl, followed by incubation overnight at 25°C for axenization. The following day, after centrifuging at 1,500 rpm for 5 min, the resulting supernatants were removed, and 7 ml of peptone yeast extract glucose medium was added to the remaining pellets; the suspensions were then transferred to T-25 flasks, followed by incubation at 25°C [
20].
Genomic DNA extraction
Genomic DNA was extracted from the cultured amoeba to identify the species of FLA and endogenous bacteria. The lysis buffer (SNET buffer) used to extract the DNA contained 20 mM Tris-HCl (pH 8.0, 1 M Tris-HCl; Dongin Biotech, Korea), 5 mM EDTA (pH 8.0, 0.5 M EDTA; Dongin Biotech), 400 mM NaCl (Duksan Pure Chemical, Ansan, Korea), 1% SDS (BiosesangTM, Yongin, Korea), and distilled water (DW), which was filtered before use using a 50-ml syringe and a 0.45-μm syringe filter (Sartorius Stedim Biotech, Göttingen, Germany). After extracting the amoebae from the 1.5% agar gels using 1× PBS, the suspensions were centrifuged at 15,000 rpm for 10 min. Following the removal of the supernatants, the remaining pellets were resuspended in protease K-supplemented SNET buffer at a concentration of 0.1 g/4 ml. The samples were incubated overnight in a 55°C-incubator using a rocker. The following day, an equal volume of phenol: chloroform: isoamyl alcohol (25:24:1) reagent was added to the samples, followed by incubation on a rocker for 30 min at room temperature. The mixtures were centrifuged at 15,000 rpm for 5 min at room temperature, and the resulting supernatants were transferred to fresh Eppendorf tubes. An equal volume of isopropanol was added, and the samples were centrifuged at 8,000 rpm for 15 min at 4°C. The resulting supernatants were discarded, and the pellets were washed with 1 ml of 70% ethanol by centrifuging at 8,000 rpm for 5 min at 4°C. The ethanol was discarded, following which the pellets were air-dried for 20 min and dissolved in diluted 1×TE buffer (10×TE buffer; Dongin Biotech).
Genetic characterization
Polymerase chain reaction (PCR) was performed to identify the amoebae isolates using reaction systems containing 2 μg of template DNA, 0.5 μl of Taq buffer, 3 μl of 10× buffer, 2 μl of dNTPs, and 2 μl each of the forward and reverse 18S rDNA primers (V4-1F 5′-GCGGTAATTCCAGCTC-3′ and V4-4R 5′-GCCMTTCCGTCAATTCC-3′) [
21]. The amplifications were performed with forward and reverse 16s rDNA primers (27F 5′-AGAGTTTGATCCTGGCTCAG-3′ and 1492R 5′-GGTTACCTTGTTACGACTT-3′) to identify the bacterial endosymbiont, and the total volume was adjusted to 30 μl using DW. The PCR conditions were as follows: initial denaturation at 95°C for 5 min, followed by 34 cycles of denaturation at 95°C for 30 sec, annealing at 55°C for 30 sec, extension at 72°C for 45 sec, and a final extension step at 72°C for 5 min, after which the samples were held at 4°C. The amplified products obtained were loaded onto 1% agar gels and subsequently sequenced commercially by Macrogene (Seoul, Korea).
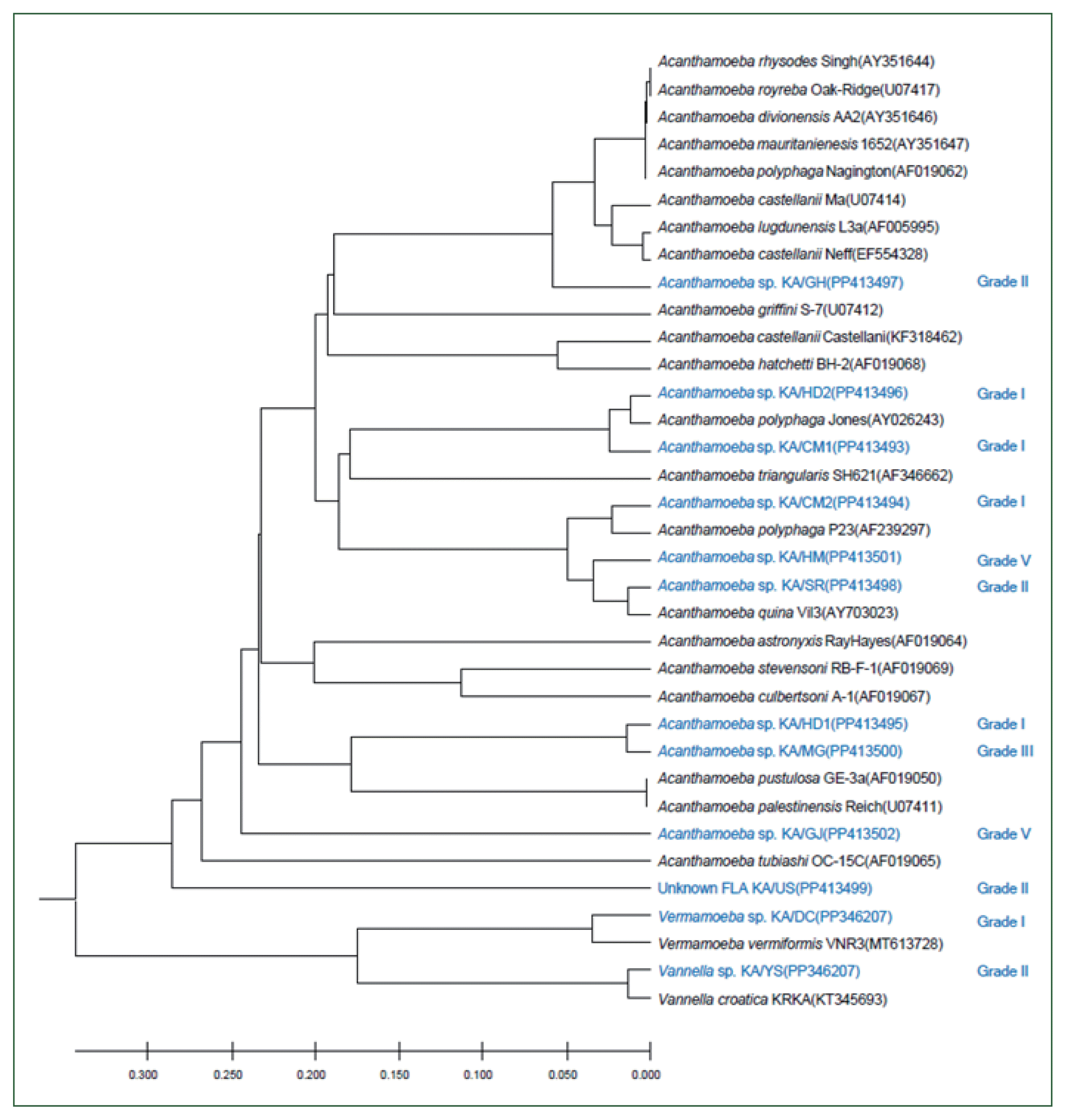
Phylogenetic analyses
The nucleotide sequences were compared with those of published strains via BLAST searches. Clustal X was used for pairwise alignment and to calculate the percent sequence identities. The percentage similarity corresponds to the number of identical sites between 2 isolates aligned against each other, as in the master alignment. The phylogeny was determined using the Clustal W2 program (
http://www.ebi.ac.uk/Tools/clustalw2/index.html) with a low gap penalty, and phylogenetic trees were constructed using TreeView software. All sequences obtained in this study have been registered in GenBank.
Transmission electron microscopy
Samples were prepared for transmission electron microscopy by initially prefixing with 2.5% glutaraldehyde at 4°C, in a phosphate buffer (pH 7.2) and postfixing with 1% osmium tetroxide using the same buffer. The material was then dehydrated through a graded series of ethanol and embedded in epoxy resin (Epon 812 mixture). Sections (1 μm thick) were stained with 1% toluidine blue for examination under a light microscope. Thin sections (50–60 nm) were prepared using an ultramicrotome (EM UC7; Leica) and double-stained with uranyl acetate and lead citrate. The stained sections were observed under a JEM-1200EXII transmission electron microscope (JEOL).
Discussion
The sustainability of FLA as bioindicators of water quality was assessed in this study. FLA from freshwater habitats characterized by varying qualities of water, ranging from Grade I to VI, were collected. As anticipated, FLA were detected in all the collected freshwater samples. Although we could not detect any significant differences among the species of FLA isolates based on the quality of the source water, the time to detection tended to shorten with the deterioration in water quality from Grade I to V. This phenomenon may indicate the higher nutrient content in more polluted water, which supports the larger populations of FLA [
22].
As shown in
Table 1, the number of coliform bacteria in the water samples increased with the decline in water quality. The poor-quality water may serve as a source of food for FLA inhabiting these habitats. The method used in the current study could serve as an additional approach for determining the water quality in different types of streams and rivers. The numbers of coliform bacteria were differences when comparing the water quality grades each time a sample collection (data not shown), indicating that numerous different approaches based on different criteria can be used to determine the water quality [
2].
A notable finding of this study is that, although FLA were detected in all the collected water samples, we isolated only 12 distinct types (
Table 1). In poor quality water (≥Grade III), obtaining monoxenic and axenic cultures of FLA is particularly difficult because of the heavy fungal and bacterial contamination. Thus, it is difficult to ascertain whether the inability to isolate FLA from Grade III and IV water reflected the effect of water quality on the presence of amoebae in this study. Nevertheless, among the FLA isolates, we discovered a species of
Vannella croatica, which, to the best of our knowledge, has not been reported from riverine waters in Korea. The genetic characteristics of this isolate (KA/YS) indicate that it is related to
Vannella croatica, the morphology and phylogenetic characteristics of which were first reported in Croatia in 2016 [
23]. However, only the trophozoite form of this species was detected; no cysts were observed in the current study. The size of the trophozoites ranged from 30 to 50 μm, and their movement was notably more rapid than that of the
Acanthamoeba species.
Microscopic observations of the FLA morphology were conducted, and the 18S rDNA genes were sequenced from the genomic DNA to identify the FLA at the species level [
4,
20]. The majority of the isolated FLA belonged to species in the genus
Acanthamoeba. Morphological analysis indicated that they were group I- to III-type
Acanthamoeba, with most having group II characteristics (
Supplementary Fig. S1). However, given that the shape of
Acanthamoeba can vary depending on the culture conditions, morphology-based assignments can prove unreliable [
4]. In the present study, amoeba with genetic characteristics similar to those of
Vermamoeba vermiformis were isolated from water samples of Grade I quality. Lee et al. [
24] indicated that, in Korea, 12.9% of tap water from buildings with storage tanks was contaminated with FLA, and the rate of contamination in highway service areas was as high as 33.3% [
24]. Nearly all the FLA detected by the authors were genetically similar to
Vermamoeba vermiformis (previously classified as
Hartmannella vermiformis) [
24]. The KFA5 isolate could represent a new allergen; the airway resistance values for this isolate were significantly elevated after 6 intranasal treatments, similar to values obtained following the administration of
Acanthamoeba [
25].
Furthermore, 16S rDNA PCR analyses of all axenic cultured isolates, except for the KA/CM1 isolate, were performed to determine the existence of endogenous bacteria in FLA. Endogenous bacteria were detected in all FLA, among which 4 species (Holosporaceae bacterium, Sinorickettsia chlamys, Candidatus Amoebophilus asiaticus, and Candidatus Paracaedibacter acanthamoebae) were identified. Based on these findings, it was impossible to discern any clear association between the endogenous bacteria in FLA and the quality of water from which the amoebae were isolated. Although genetically distinct, certain amoeba isolates (KA/US, KA/CM2, and KA/MG) hosted a similar complement of symbiotic bacteria. In contrast, genetically similar FLA (KA/HD1 and KA/MG) contained notably different symbiotic bacteria. Interestingly, although large numbers of Acanthamoeba previously isolated from tap water have been shown to lack bacterial endosymbionts, almost all amoebae isolated from river water were found to harbor symbiotic bacteria in this study. These findings indicate that the endosymbiotic associations between Acanthamoeba and bacteria can appear and disappear. In the present study, endosymbiotic bacteria were detected in the KA/GJ isolate based on the 16S rDNA PCR analysis but were not visible under the electron microscope.
Species in the
Holosporales order, which are an alphaproteobacterial lineage encompassing bacteria obligatorily associated with multiple diverse eukaryotes within isolated FLA [
26], were also detected in the current study. These bacteria are phylogenetically related to the symbionts of other ciliates and diplonemids, forming a putatively rapidly evolving clade within the
Holosporaceae family; the characteristic features of this clade include the presence of specific secretion systems, and they are speculated to have a mild parasitic effect on their hosts [
26]. The endosymbiont identified in the
Acanthamoeba KA/SR was found to be very similar to that detected in KA/E9 isolated from humans with keratitis [
27]. However, there is no strong evidence to indicate that this endosymbiont plays a pathogenic role. Interestingly,
Sinorickettsia chlamys (GenBank No. AY174894), previously identified as an intracellular proteobacterium in the Chinese scallop
Chlamys farreri, a species of marine bivalve, was identified in the
Acanthamoeba isolate KA/SR in this study. These bacteria are phagocytosed by different aquatic animals and subsequently persist as intracellular “organelles.” However, the roles played by these bacteria in animal cells have not been established and warrant further research.
WQI is a simple tool that assigns a single value to water quality by considering a certain number of biological, chemical, and physical parameters (also called variables) to represent the water quality in an easy and understandable way [
28,
29]. Despite the efforts of researchers around the world, an index that can be universally applied by water agencies, users, and managers in different countries is lacking [
30]. In the current study, we aimed to use FLA as an indicator of water quality. However, it takes a long time to isolate and culture amoeba, and using them with the currently available methods is challenging. Therefore, it is necessary to develop a simple kit by developing the PCR and Loop-Mediated Isothermal Amplication methods that can easily detect FLA in water samples.
In conclusion, although we were unable to provide conclusive evidence indicating an association between FLA (or their endosymbionts) and the quality of water from which they were isolated, we did detect significant increases in the sizes of the amoeba populations with the deterioration in the water quality. This result showed that eutrophication of fresh water is positively related to the proliferation of free-living amoeba.