Introduction
Chromosomal DNAs in eukaryotic cells are present in a highly organized structure by being packaged into nucleosomes, the structural unit of chromatin, in combination with histone proteins [
1]. Together with the histone globular domains, the unstructured side chains of specific amino acid (aa) residues provide binding platforms for various chromatin-binding proteins including chromatin remodelers, DNA/histone-modifying enzymes, and general transcription factors [
2]. The physicochemical nature of globular domains and side chains profoundly affect the macroscopic chromatin structure and gene expression by modulating attachments of structural/functional proteins involved in the biological processes. The nucleosomal histones are modified by adding a wide variety of chemical groups such as methyl, acetyl, phosphoryl, propionyl, serotonyl, and acyl groups, to meet the essential biological demands [
2,
3]. Mis-regulated histone modifications may result in inappropriate activation or repression of various genes, which can lead to developmental anomalies and/or pathobiological progression of various diseases including cancer [
4].
Epigenetics refers to the heritable modification of gene expression without any change in DNA sequences, thereby introducing diverse phenotypes among cells with identical genetic backgrounds in multicellular organisms. The chromosomal events are also implicated in maintaining the integrity and stability of gene expression profiles in the resulting variants [
5,
6]. During the epigenetic processes, gene expression is regulated at various levels in response to the DNA and histone modifications, as well as the actions of non-coding micro RNAs [
5]. Of the diverse histone modifications, methylation and acetylation are the most well studied. Reversible addition or removal of methyl and acetyl groups induces a structural shift in chromatin from compact fibers to relaxed ones, or vice versa. These conformational alterations are closely related to the modulation of the frequency, activity, or extent of nearby gene expressions by influencing the accessibility of the transcriptional machinery [
2].
A series of methylated Lys residues associated with the epigenetic mechanisms have been reported in nucleosomal histones, such as mono (me1), di (me2), and tri (me3)-methylation of Lys residues at positions 4, 9, 27, 36, and 79 of histone H3 (H3K4, H3K9, H3K27, H3K36, and H3K79); at positions 20 and 59 of histone H4 (H4K20 and H4K59); and at position 26 of histone H1B [
2,
7]. The methyl groups are transferred from S-adenosyl methionine (SAM) to the ε-amino group of Lys, and the biochemical reactions are catalyzed by a group of enzymes, collectively termed histone Lys methyltransferase (HKMT) [
8]. Except for the Disruptor of Telomeric Silencing1 (DOT1) family, which is responsible for the methylation of H3K79, almost all HKMT proteins possess a relatively conserved domain, the suppressor of variegation (Su(var)3–9), enhancer of zeste (E(z)), and trithorax (SET) [
7]. The methylated histones may specify both active and repressed chromatin states, depending on the positions of modified Lys residues and the degree of methylation. However, the biological role of each HKMT remains elusive [
2,
7]. In general, H3K4me3, H3K79me2, and H4K20me1 are involved in transcriptional activation, whereas H3K9me2, H3K9me3, and H3K27me3 are associated with gene silencing [
2,
9].
Protein genes involved in epigenetic processes, such as those encoding DNA methyltransferase (DNMT) and HKMT, show punctate distribution patterns across the genomes of eukaryotic taxa [
10,
11]. This suggests that eukaryotic organisms have uniquely evolved epigenetic toolkits to meet individual requirements for adapting to different environmental conditions and/or regulating germline/soma differentiation. Defense against mobile genetic elements provides an additional driving force for the mosaic evolutionary pathway. The presence and biological implications of the epigenetic processes, or at least methylated cytosine-mediated gene silencing mechanism, have been highly suspected in lower animals, including those of the phylum Platyhelminthes, owing to the lack of DNMT1 (maintenance of cytosine methylome) and DNMT3 (de novo generation of cytosine methylome)-like enzymes [
10–
12]. However, recent studies have shown that biochemical modifications of DNA and histones are involved in the spatiotemporal regulation of gene expression without genetic alterations in platyhelminth species [
13,
14].
Clonorchis sinensis belonging to the class Trematoda of the phylum Platyhelminthes parasitizes the biliary tracts and causes clonorchiasis in humans. Clonorchiasis poses a great socio-economic burden in endemic areas, including China, Korea, and Vietnam [
15]. Humans are accidentally affected by the parasitic disease by consuming raw or undercooked freshwater fish containing
C. sinensis metacercariae. The metacercariae excyst in the duodenum and grow into mature adults in the bile ducts. Patients with clonorchiasis exhibit a wide spectrum of clinical symptoms, ranging from obstructive cholangiopathy to cholangiocarcinoma, depending on the rate and duration of infection [
16]. The hepatobiliary worm is categorized as Group I biocarcinogens by the World Health Organization [
17]. Similar to other digenean parasites,
C. sinensis has a complex life cycle and undergoes metamorphosis in its respective hosts under different biophysicochemical environments: 4 stages from miracidia to cercaria in snail, the metacercaria stage in freshwater fish, and the juvenile/adult stage in mammal [
18]. Furthermore, approximately 32.0% of the parasite genome comprises diverse repetitive elements, including long terminal repeat (LTR) and non-LTR retrotransposons [
19,
20]. In addition to the parasitic life mode, this genomic feature implies that certain types of molecular mechanisms operate effectively in
C. sinensis for regulating gene expression in a timely manner and/or defensing against the selfish intragenomic invaders [
10]. Reversible modification of the chromatin structure induced by methylated DNA and/or histones may be a strong candidate for the epigenetic gene regulation. We have previously reported the structural and biochemical properties of a DNMT homolog in
C. sinensis [
21]. In this study, we analyzed HKMT proteins by screening the entire
C. sinensis genome. Temporal expression patterns of these HKMT genes and the effects of methyltransferase inhibitors on the expression of several functional genes were also investigated in the liver fluke to determine their role as epigenetic regulators.
Discussion
Since DNMT2 was shown to be an RNA methyltransferase [
30] and platyhelminths belong to the so-called DNMT2-only organisms [
10], the presence of DNA methylation-related epigenetic machinery and/or its practical implications in gene regulation have been highly disputable for a long time in the lower animals. However, recent studies have provided multiple evidences revealing that the epigenetic machinery effectively operates in the lower animal group [
14,
21,
31]. In addition to the previously described 5-methylcytosines [
21], this study showed that methylated histone variants are intimately involved in the epigenetic regulation of
C. sinensis genes. For the differential histone methylation, genes encoding most of the HKMT family members, which have been widely identified across animal taxa, are preserved in the liver fluke and other platyhelminth genomes, while some genes were specifically expanded in a series of donor organisms. The temporal expression profiles of
CsHKMT and other functional genes suggest that histone methylation is deeply involved in epigenetic pathways that contribute to age-associated phenotypes, such as reduced fecundity in
C. sinensis. This aging process may be reversed by inhibiting CsHKMT activities.
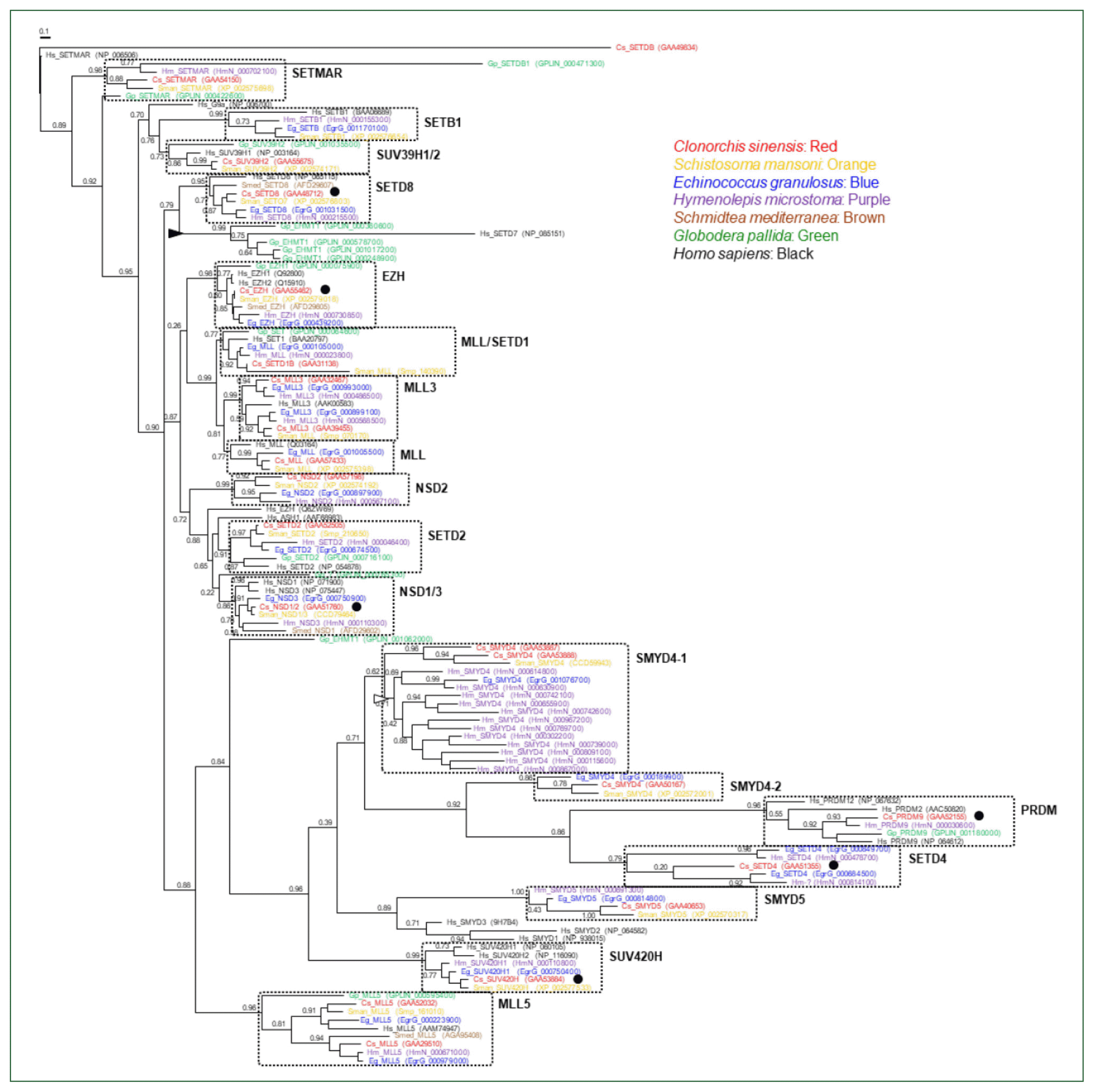
The mammalian SET-HKMT family can be categorized into approximately 6 subfamilies (SET1 for H3K4, SUV39 for H3K9, SET2 for H3K36, EZH for H3K27, SMYD for H3K4, and PRDM for H3K9) and an extra group comprising multiple heterogeneous proteins based on structural and enzymatic characteristics [
22]. Most of these subfamily members have specific orthologous and/or biochemical counterparts in plants [
32], demonstrating that the protein family may have varied during an early stage of metazoan evolution before divergence of animals and plants. Parasitic helminths examined in this study have well preserved all these subfamilies: SETD1, MLL, and MLL3 lineages in SET1; SETMAR, SETB1, and SUV39H1/2 lineages in SUV39; NSD1-3 and SETD2 lineages in SET 2; EZH lineage in EZH; SMYD4/5 lineages in SMYD; PRDM lineage in PRDM; and the other heterogeneous SET4/8, SUV420H, and MLL5 lineages (
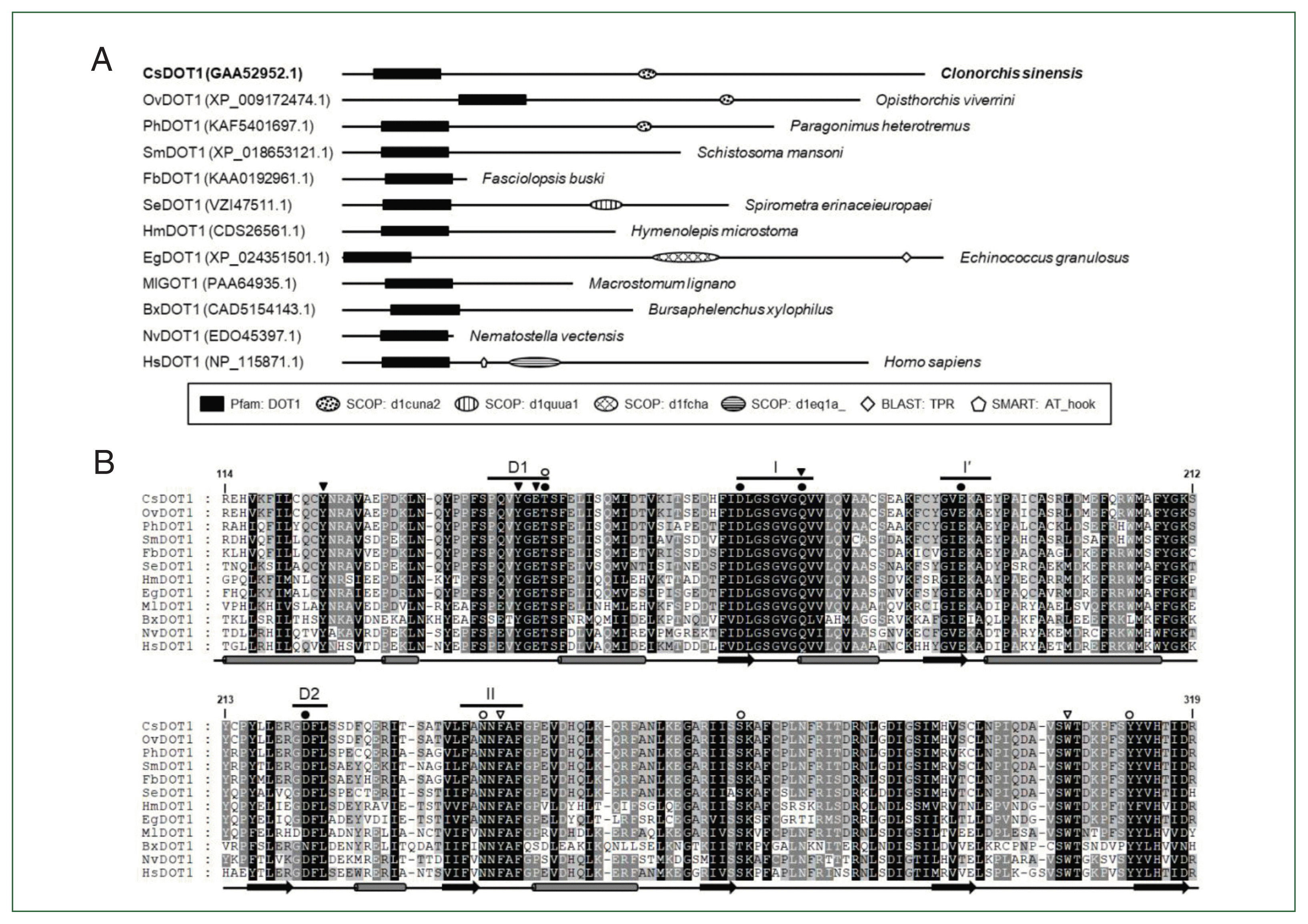
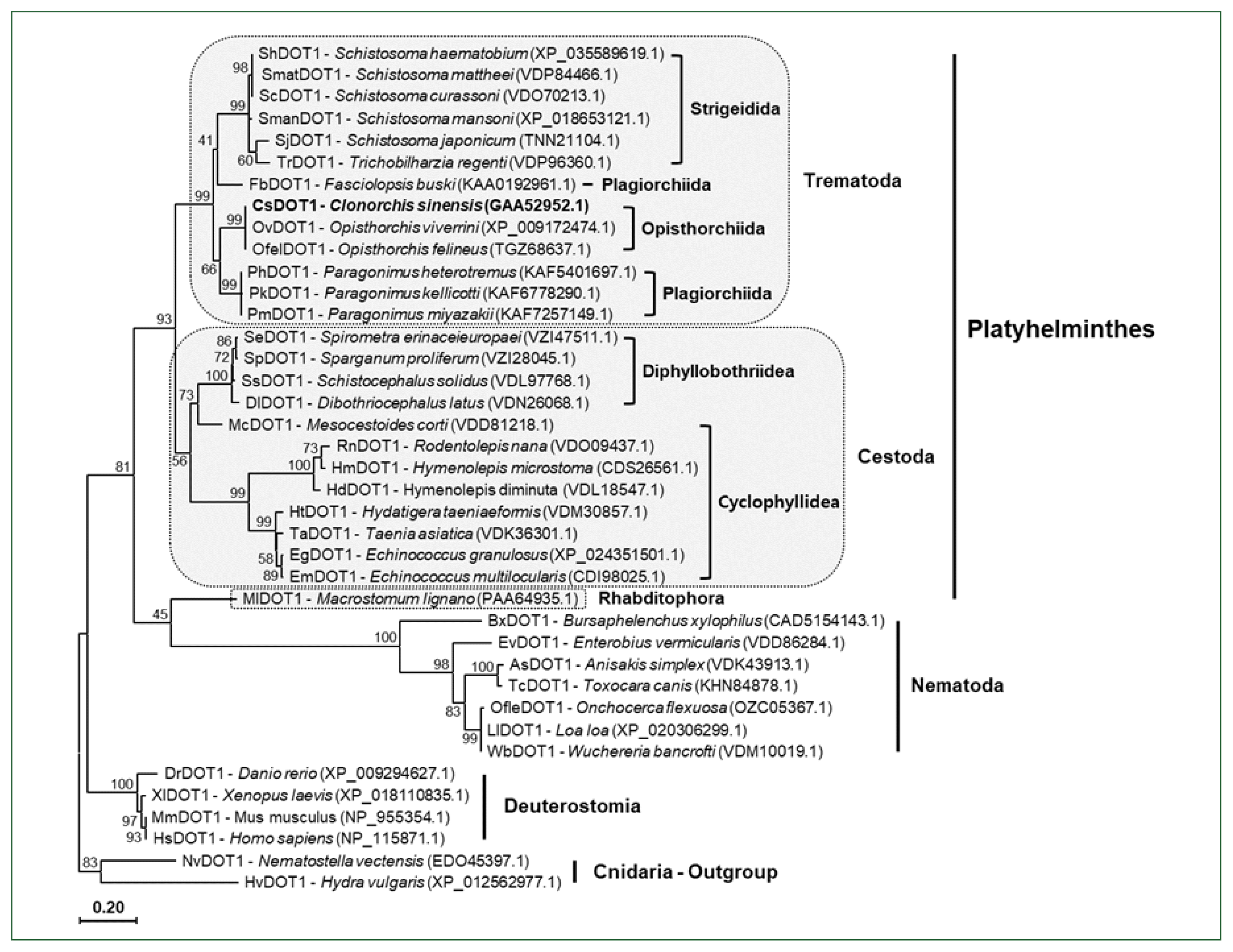
Fig. 2). A single gene encoding the non-SET HKMT protein DOT1, which is responsible for H3K79 methylation, was also well conserved in each parasite genome (
Figs. 3,
4). The high gene dosages were in contrast to that of
DNMT gene. Platyhelminth genomes preserved only a single
DNMT2 homolog but no DNMT1 or DNMT3 homologs [
21].
The life cycle of
C. sinensis comprises 7 distinct developmental stages involving 3 sequential hosts: egg; miracidium, sporocyst, redia, and cercaria in freshwater snail; metacercaria in freshwater fish; and adult in mammal [
33]. The worms at discrete stages exhibit great differences in morphology, size, structure, and tissue/organ composition. Therefore, it can be postulated that the parasite is highly dependent on methylated histone-mediated epigenetic processes for stage-specific gene expression profiles on an identical genetic background, as has recently been examined in
S. mansoni [
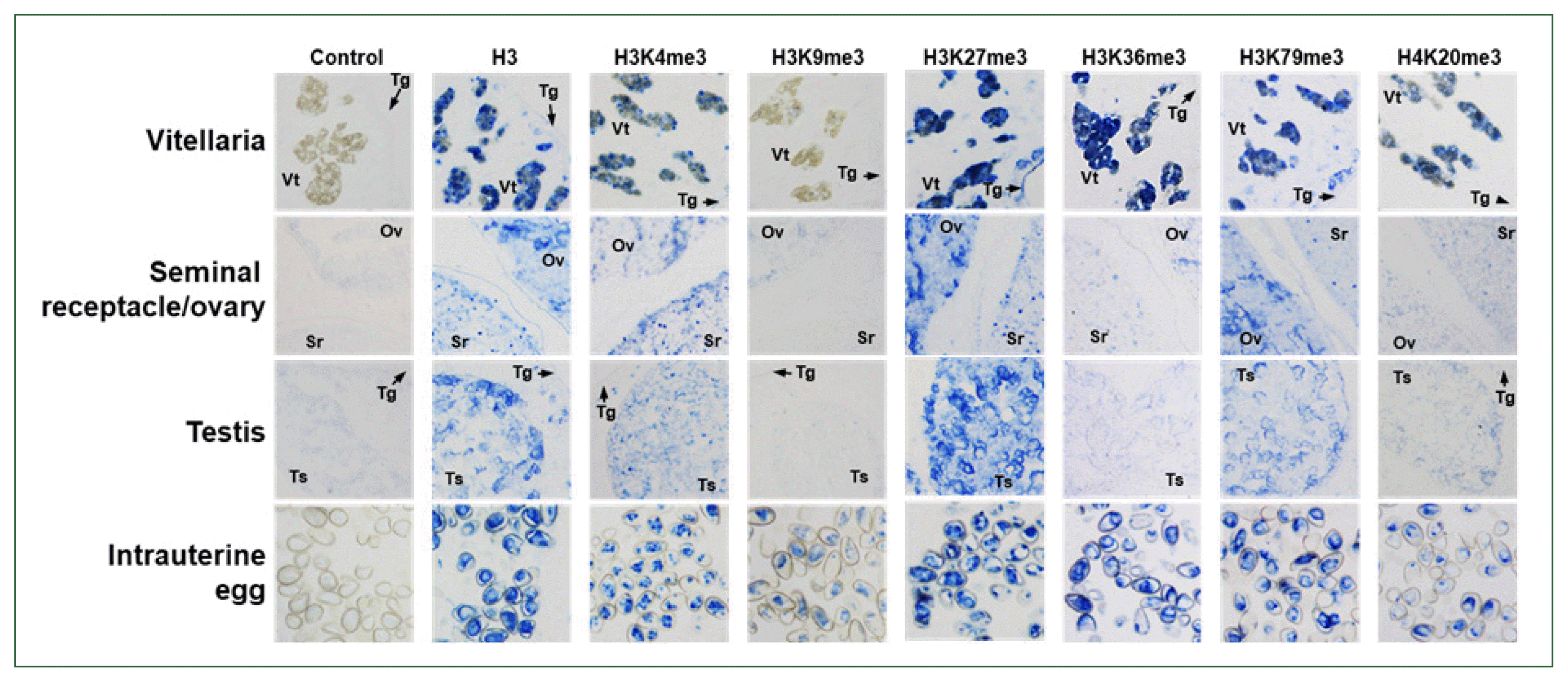
14]. Consistent with the large enzymatic pool, diversely methylated histones were identified in whole-worm sections of
C. sinensis adults (
Fig. 5). In general, the methylated forms of H3K4 and H3K79 are associated with the transcriptional activation of relevant genes, whereas those of H3K9, H3K27, and H3K36 are closely linked to euchromatic and/or heterochromatic silencing. In contrast to the mono- and di-methylated forms, the tri- methylated form of H4K20 also serves as a silencing marker [
7]. The relative densities of these epigenetic markers were substantially different among the
C. sinensis tissues/organs. H3K79me3 and H3K27me3 are likely the major transcriptional activator and repressor, respectively, in the vitellaria, ovaries, and testes. The repressive H3K36me3 was also highly enriched in the vitellaria. The activating H3K4me3 and repressive H4K20me3 were moderately detected in these organs. However, all histone markers examined in this study were expressed at relatively high levels in intrauterine eggs (
Fig. 5).
The genome-wide methylation status of DNAs and histones is significantly altered by reversible demethylation-methylation events throughout the development and organogenesis in animals [
34]. The major epigenetic markers, especially H3K4me3 and H3K27me3, also fluctuate highly in
S. mansoni in accordance with developmental progress [
14]. Although the methylome changes were not investigated following the development in
C. sinensis in this study, it was apparent that the genome-wide densities of methylated histones in intrauterine eggs differ from those in other tissues/organs in
C. sinensis adults. The fertilized ovum (i.e., zygote) of
C. sinensis and
S. mansoni is enclosed in a proteinaceous shell within the ootype and embryonate into the miracidium while passing through the uterine tube. Finally, fully embryonated egg is released from the uterine pore and hatches in fresh water (
S. mansoni) or in the first snail host (
C. sinensis) [
35]. The oscillating genome-wide frequency of H3K27me3 during schistosome development was the lowest at the miracidium stage, probably due to global demethylation after the fusion of gamete cells [
14]. However, the densities of the diversely methylated histones, including H3K27me3, were considerable in the embryonated eggs of
C. sinensis. Considering that the genome-wide frequencies of methylated histones increase according to the maturation and/or aging of donor organisms [
34], this observation may reflect differentially operating epigenetics between the 2 trematode parasites. Alternatively, this fact possibly results from different degrees of maturation, i.e., developing vs. fully developed miracidia. Further investigation on the different maturation stages of miracidium may help address this issue.
The fertility rate of the hermaphroditic
C. sinensis gradually declines to a very low level as the worm ages in the definitive hosts after reaching a peak [
36]. The temporal expressions of
Clonorchis genes, such as
CsTyrs and
CsMbs, the protein products of which play key roles in sexual reproduction, appears to be synchronized with the change in fecundity [
25,
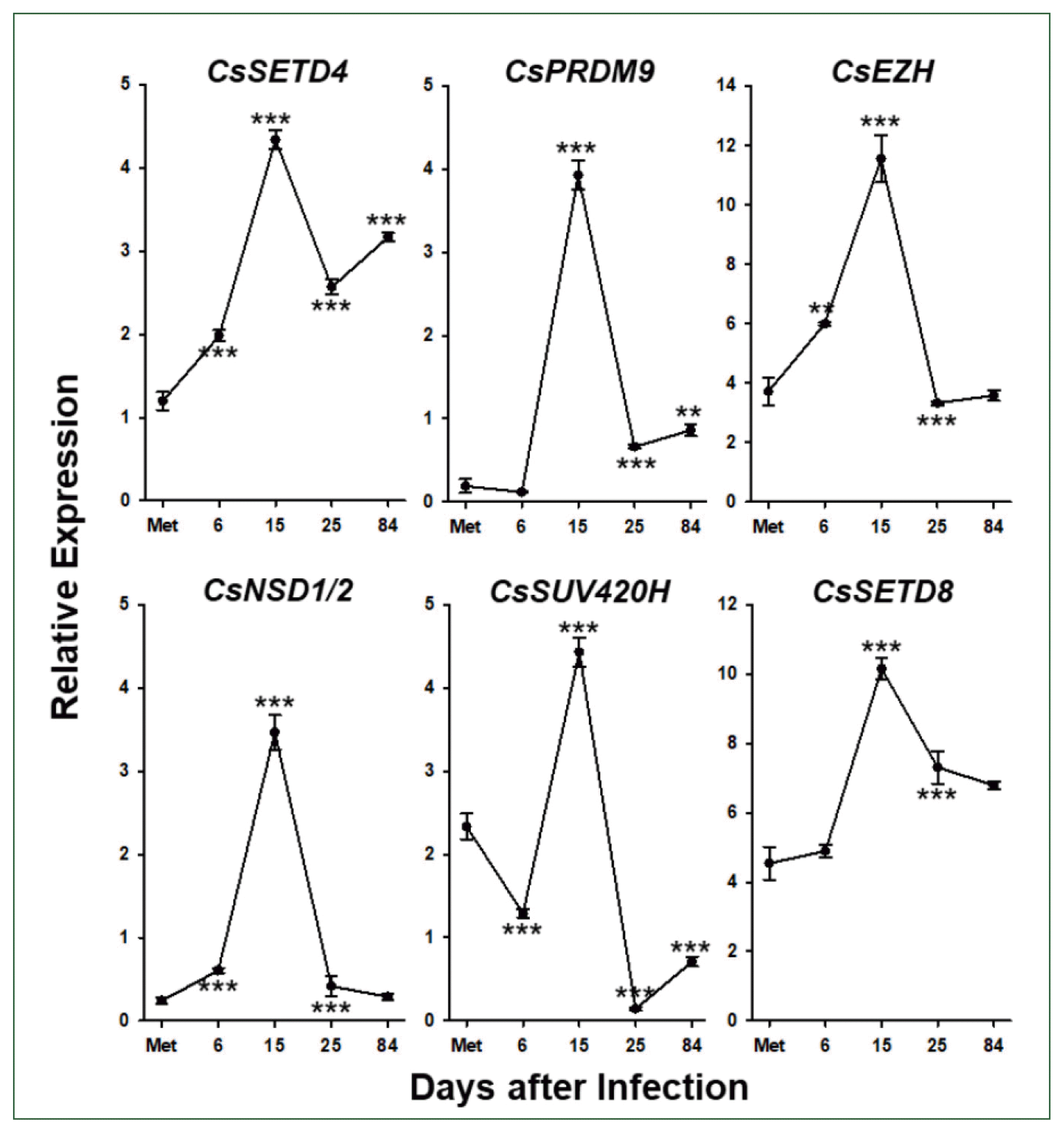
29]. However, the expression of genes encoding epigenetic writers (HKMTs and DNMT) and readers (MBDs) significantly increased in later adulthood (
Fig. 6 and
Supplementary Fig. S3). Therefore, it could be postulated that the fecundity reduction is attributable to the down-regulation of essential genes corresponding to the increasing frequencies of epigenetic markers. This presumption was, at least in part, supported by observations showing that functional gene expressions are differentially regulated by histone/DNA methylation depending on the age of
C. sinensis adults (
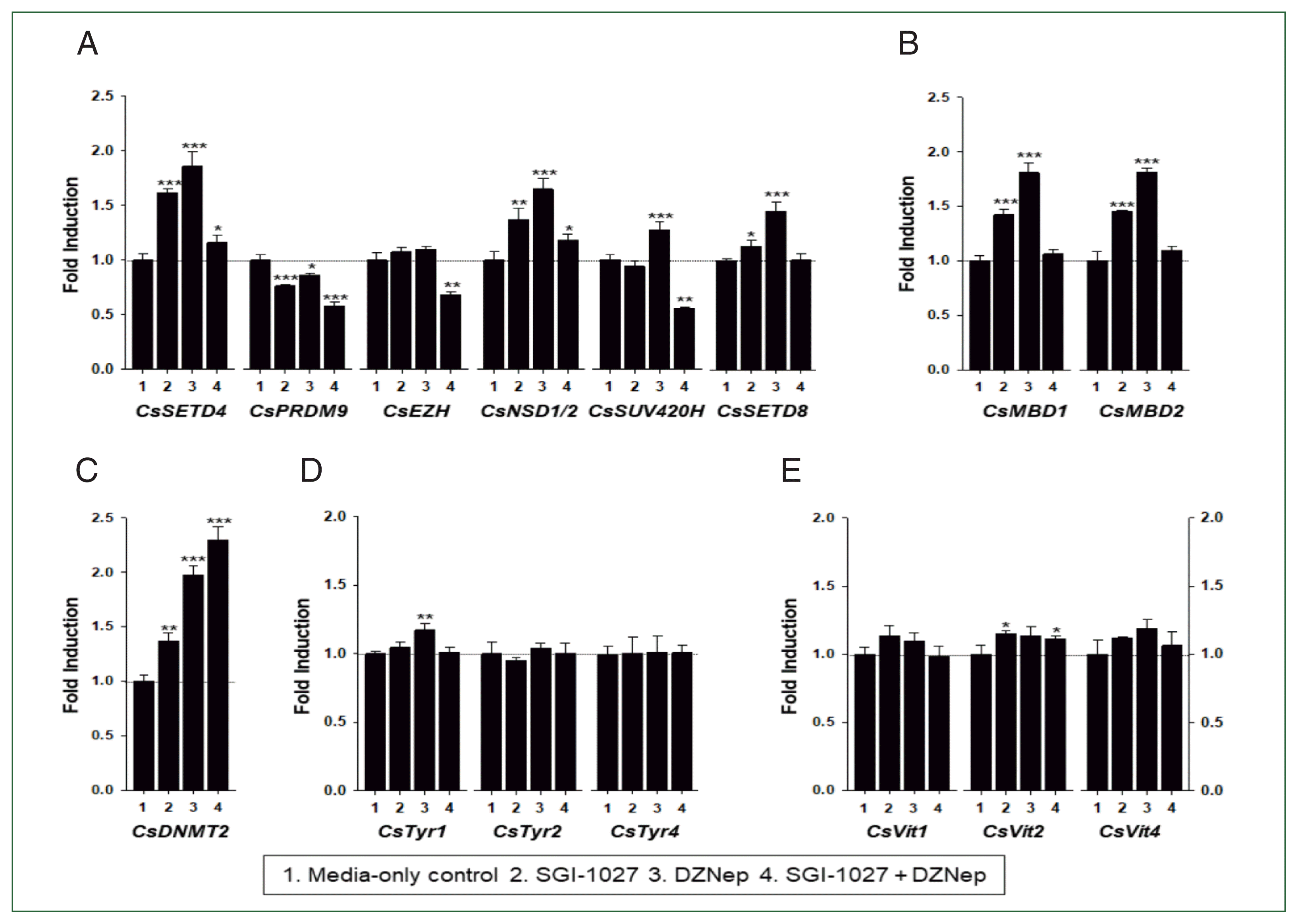
Figs. 7,
8). In 42-day-old worms, epigenetic tool genes were likely repressed by DNA and histone methylation, but those related to eggshell formation were not under the epigenetic control (
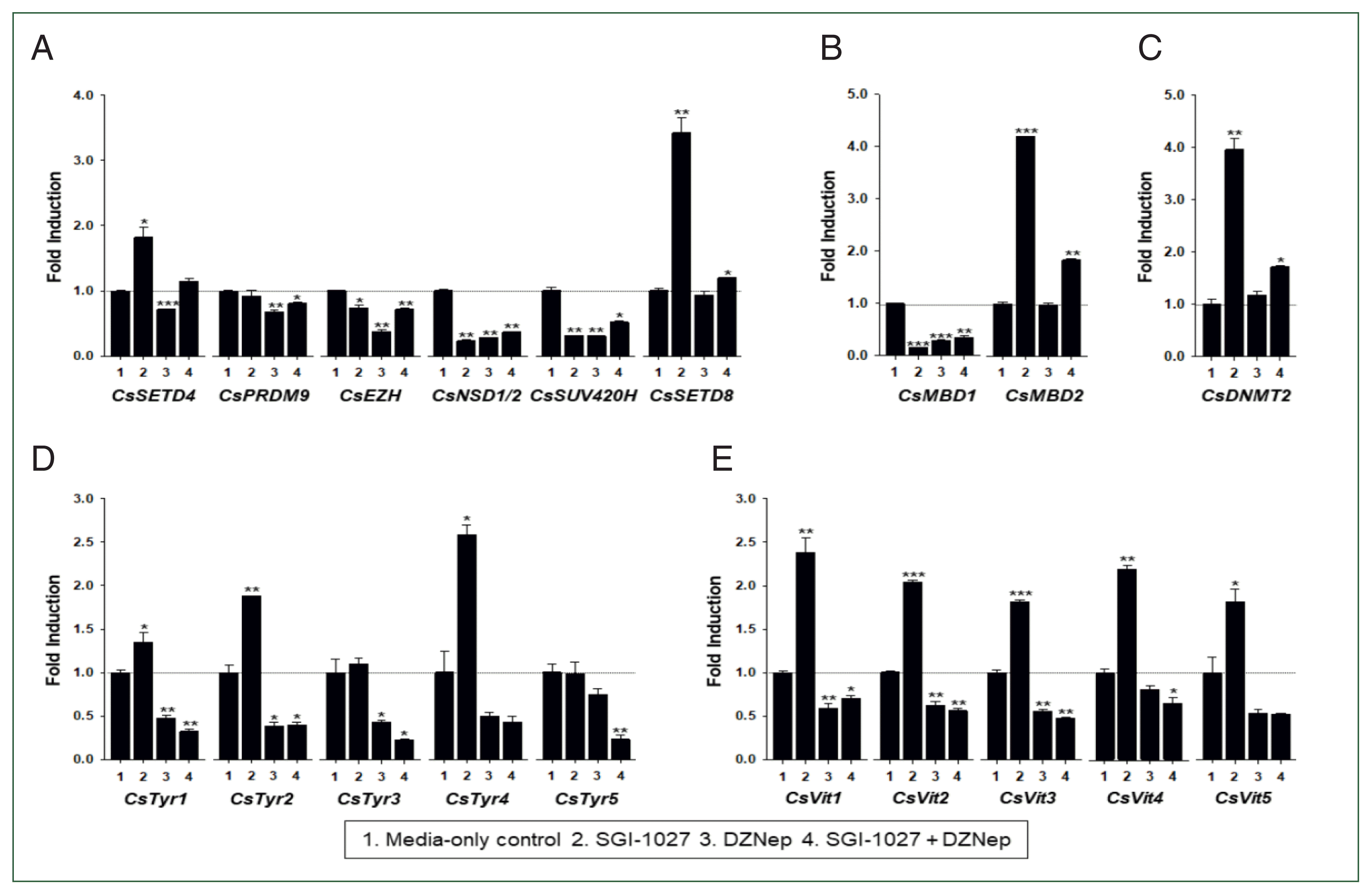
Fig. 8). In contrast, eggshell-related genes and some epigenetic tool genes were epigenetically silenced mostly by DNA methylation in 140-day-old worms. The significant down-regulation of
HKMT and
MBD1 by treatment with methyltransferase inhibitors in the older worms further revealed that epigenetic activators, such as H3K4me3, are intimately involved in the up-regulation of epigenetic tool genes (
Fig. 7). S-adenosylhomocysteine (SAH) hydrolase-inhibiting DZNep and SAM-competitive SGI-1027 are known to be specific to EZH2 [
37] and DNMTs [
38], respectively. However, these molecules may act as pan-methyltransferase inhibitors, as most cellular methyltransferases use SAM as a methyl donor and are inhibited by SAH, which is generated by the methylation reaction and hydrolyzed into adenosine and homocysteine by SAH hydrolase [
39]. Therefore, the modulated gene expressions by the chemical treatments are likely the sum of broad methyltransferase inhibition rather than restriction to a specific CsEZH or CsDNMT2 blockade. Nevertheless, our data clearly demonstrate the significant role of the epigenetic machinery in the temporal regulation of
C. sinensis genes, especially those involved in sexual reproduction.
Methylated DNA and histones are highly interrelated and depend mechanistically on each other for the epigenetic regulation of gene expression and/or maintenance of chromosomal integrity [
40]. Most multicellular eukaryotes express 3 DNMTs that generate (DNMT3a and DNMT3b) and maintain (DNMT1) the genome-wide 5mC methylomes. However, the genomes of
C. sinensis and other platyhelminths encode only a single
DNMT protein homologous to the mammalian DNMT2 [
21] and the DNMT2-like protein seems to provide enzymatic activity responsible for the generation and/or maintenance of DNA methylomes [
21,
31]. This situation is highly contrasted to the well-preserved
HKMT dosages in these invertebrates (
Figs. 2,
4). The reduced dosage of
DNMT gene would directly contribute to the low frequency of genome-wide 5mC in
C. sinensis and
S. mansoni [
21,
31]. However, whether the imbalanced genetic dosages of
DNMT and
HKMT in the DNMT2-only organisms influence on taking initiative during epigenetic process remains unclear. Locus-specific methylome maps previously obtained in
S. mansoni were constructed using chromatins extracted from whole worm bodies [
14,
31]. Cells comprising various tissues/organs in multicellular organisms are heterogeneous in terms of epigenetic modifications [
34]. Consequently, cell- and tissue-specific methylome patterns and their implications in the regulation of nearby genes in parasitic trematodes have not yet been defined. Future investigations with homogeneous cell populations should address issues regarding the detailed interrelationship between DNA and histone methylation, and the tissue-specific methylome construction in association with the development and/or maturation stages in the DNMT2-only organisms.